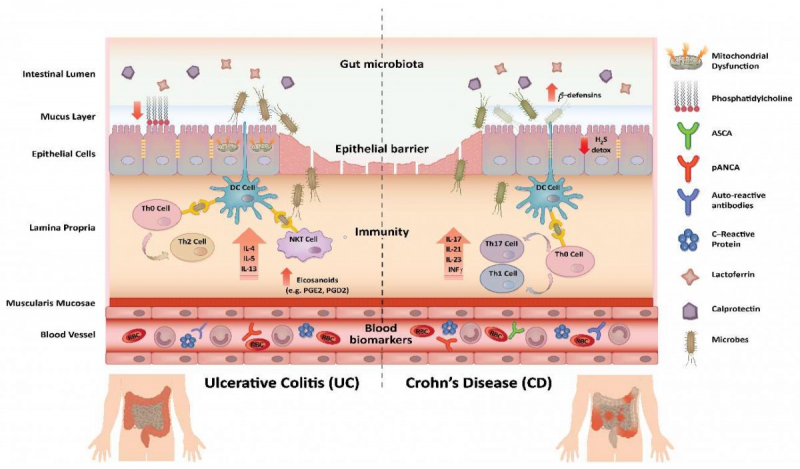
Millions of people all over the world suffer from various inflammatory bowel disorders. These are severe chronic illnesses that are associated with recurrent pains, diarrhea, weight loss and even fever. As of today, the two most widespread are ulcerative colitis and Crohn's disease. For one, every 200th citizen of Norway has ulcerative colitis, and Crohn's disease is diagnosed in almost every 300th German.
The clinical picture of different inflammatory bowel disorders can be similar, so it is essential for the doctors to have reliable diagnostics methods that make it possible to identify what exactly a person suffers from and what treatment they need. Among the promising means is a metagenomic analysis of bowel microbiota that allows for telling which useful and harmful microorganisms live in a patient’s gastrointestinal tract. The Metagenomics diagnosis for inflammatory bowel disease challenge (MEDIC) international competition for bioinformatists that took place in 2019-2020 was dedicated to studying this method’s potential.
A team that included ITMO student Artyom Ivanov and his research advisor, the head of the international laboratory "Computer Technologies" Vladimir Ulyantsev succeeded in getting first place in two of the competition’s categories.

From the USA to Japan
The competition took place online; in order to participate, you had to register at the organizer’s website before the middle of January 2020.
“Last fall, Vladimir Ulyantsev showed me a letter with a proposition to take part in this competition,” remembers Artyom Ivanov, “and I thought that competitions aimed at searching for a more precise diagnostics method is something interesting. We were already developing some programs for metagenomic microbiota analysis at our lab, so we decided to show up.”
Vladimir Ulyantsev confesses that at the time they applied, he didn’t believe that they could win. This is understandable: the competition was severe, as the event brought together 26 teams from all over the world. Ten represented Italy, Greece and Japan were represented by six teams each, and there were also three teams from Finland and three - from the USA.

Two in one
The competition’s participants were to solve two tasks, one in each category. In fact, this turned every category into a competition of its own, with specific goals, its own scores and winners.
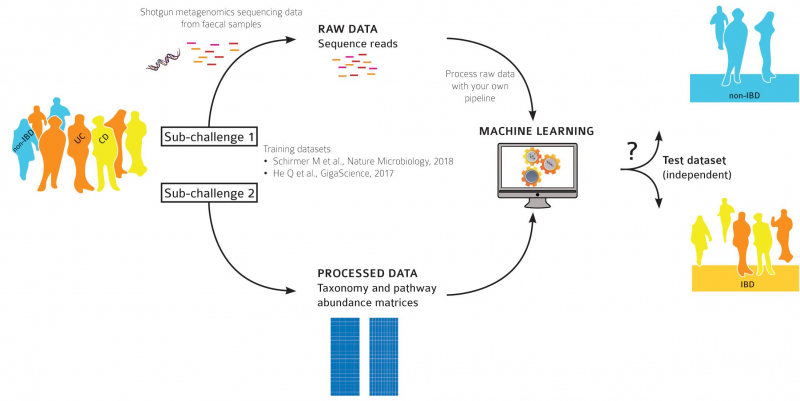
Both were organized in a standard format of machine learning tournaments: first, the participants were given tagged data of a microbiota study of 150 patients. This data was to be used for teaching a computer algorithm to identify a person’s condition: whether they are healthy, have ulcerative colitis, or Crohn's disease. After that, the analyser algorithm had to check a set of similar untagged data and make conclusions on which category each of the patients belongs to.
“In the first category, we knew which bacteria inhabit the bowels and how many of them are there,” says Artyom Ivanov. “This data didn’t call for any pre-processing, we launched our machine learning algorithms straight away. We’ve tried different boosting algorithms, and one of them worked. The second task was a lot more interesting: we only got raw data from metagenome readings, short DNA sequences from microbiota of patients about whom we knew nothing. We had to identify the features that were essential for making a diagnosis, and make predictions based on untagged data. We used our MetaFast program that identifies some related structures from the entirety of the information on the DNA, which can be the essential ones. Based on these elements, we launched machine learning and developed our classifier.”
Playing blind
MEDIC also had some features that made it different from regular machine learning competitions. Usually, teams at such competitions form a hypothesis, teach an algorithm in accordance with it, launch it on untagged data, send it to organizers and watch in real time mode how well it manages with a task and what position they have in the scoreboard. Then they get feedback which they use to improve this algorithm or develop a new one.
In MEDIC’s case, the teams got no feedback. They could do any number of attempts, but didn’t get any information on their success or their opponents’ progress.
“This was like playing blind,” comments Artyom Ivanov. “There are several ways to adjust to competing like that. One of the classical approaches is cross validation, when we leave part of the data untouched, i.e. we don’t use it to teach the algorithm but only use it to test it. Another problem is that any work with DNA requires very big data: it takes about a week to develop, teach and test an algorithm. In the course of the competition, we made about thirty attempts, but sent only the best ones.”
Another way to find the correct approach was searching for additional data. ITMO’s team succeeded in finding another set of tagged data on the internet and also use it for teaching and testing their analyser.
“Data is the most important thing in machine learning,” explains Vladimir Ulyantsev. “We can endlessly train the algorithm, but it won’t measure to anything in comparison with those by leading enterprises that teach theirs using vast amounts of data. This is why we searched for and uploaded additional data from open sources that were not provided by the organizers but fitted our tasks. This might well be the reason why we won.”
Just predicting is not enough
Apart from being a competition, MEDIC also had a scientific aspect, which is of great importance for both organizers and participants. Following the competition’s results, the winning teams are invited to work on joint publications in cited journals. As the ITMO team won in both categories, they plan to get involved in this initiative.
“The issue of diagnosing these diseases is an acute one,” stresses Artyom Ivanov. “In the course of the competition, we just identified the metagenomic features that can be used for diagnosing them. It would have been interesting to learn how and why they are related to the diseases’ development. This is how it is with just about any medical research: machine learning is good, but if we want to bring our results to some doctor, we need to explain what’s inside our black box - they can’t just blindly believe in a model.”

Difficult and interesting
Artyom Ivanov confesses that the competition has become “a big and important experience” for him.
“I’ve never worked with such a vast volume of data before,” he says. “It’s difficult and interesting: you need to understand how to include the data in the algorithm, how to process them, you need to keep track of time in order to keep up with the schedule.”
Vladimir Ulyantsev believes that the conduct of such competitions is of importance for the community of those who focus on metagenomic analysis-based research and treatment of diseases.
“As per my experience, working with diagnoses is quite hard. A diagnosis is data with a high level of noise: one doctor makes one diagnosis, another makes a different one. It’s good that they’ve started to organize such competitions, and it's great that we won this one,” concludes the scientist.