DNA machines
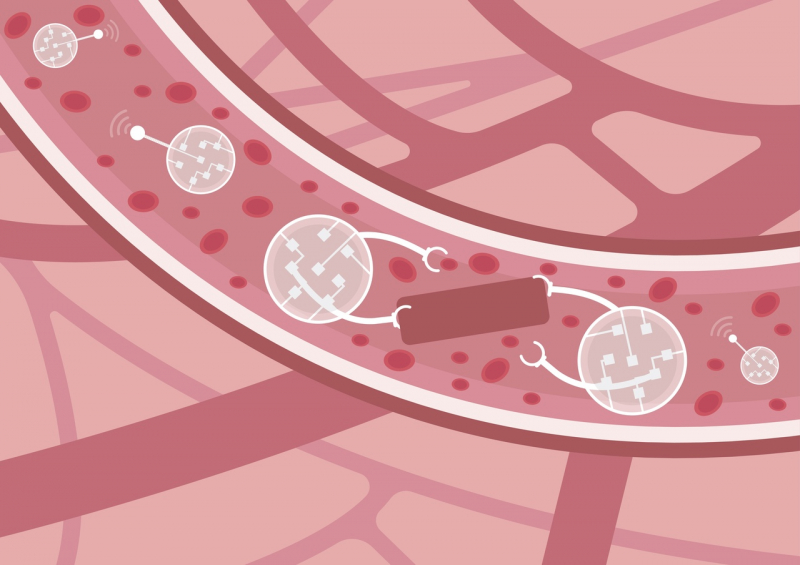
DNA robots (or DNA machines) aren’t our invention; it’s a common term for artificial molecules – intellectual systems consisting entirely of nucleic acids. Under certain conditions, such structures acquire a number of unique functions: they can locate targets (certain DNA or RNA chains), connect with them, disintegrate or change their spatial organization, block their replication, or deliver therapeutic agents to cells.
At the foundation of such artificial compounds lie deoxyribozymes, catalytic molecules with fermentative functions. They can “catch” and store magnesium and, once there is enough of it, they can disintegrate the target. Once a robot reaches a cell and determines that it’s cancerous or infectious, it joins the chain and blocks the vital genes of this cell, thus destroying it.

At the same time, there is a system to ensure the safety of healthy cells – even one nucleic replacement in the sequence-tagged site is enough to stop the robots from executing their functions. It’s important to note that DNA robots aren’t proteins, so there is no need to provide special conditions for their storage.
The same principle is used in diagnostics for typing, for example, singular nucleic replacements – a rather hard task in diagnostics.
At SCAMT, Dmitry Kolpashchikov’s laboratory Molecular Robotics and Biosensor Materials is engaged in such research. Their activities mainly lie in two fields: the use of DNA robots and DNA nanomachines in treatment of cancerous and infectious diseases and the development of DNA sensors and testing systems.
Using DNA sensors and DNA robots for diagnostics
Our team has brought together seven laboratories of the ChemBio Cluster and at the moment, we’re engaged in the development of a system for express diagnostics of infectious diseases of the central nervous system, commissioned by the Federal Medical-Biological Agency. We have already designed a working prototype of the diagnostics platform and tested it with the help of our partners: Research Institute of Infections in Children, Research Institute of Influenza, and Pasteur Research Institute of Epidemiology and Microbiology.
Infectious diseases of the central nervous system, such as meningitis and encephalitis, have high mortality rates and cause disabilities if treatment isn’t provided on time.
The main challenge that doctors face are the various limitations that make the diagnostics process too time-demanding. Nowadays, the most commonly-used express diagnostics methods are the molecular-genetic kind, but they don’t allow us to analyze the entire spectrum of possible pathogens and require special equipment and qualified specialists. Often, the samples need to be delivered to special laboratories, which is time-demanding and requires suitable transportation conditions.
Our team’s task was to develop a system that would be as fast, user-friendly, mobile, and automatized as possible so that even regular medical workers could use it. At the same time, the system must detect as wide a spectrum of pathogens – bacteria, viruses, and fungi – as possible, and work under any conditions.
Solving this task became possible thanks to the use of our sensors.
Our DNA sensor is composed of two parts: two sequences that detect a pathogen’s RNA and two halves of G-quadruplex (guanine tetrad). Once a special matrix appears in the compound – for example, in the pathogen’s DNA or RNA – the system connects the two halves and forms a G-quadruplex. This launches a sequence of catalytic reactions which eventually results in the studied sample turning brown.
Such visualization is very useful, as no additional equipment is necessary to detect pathogens. The fact that we don’t use proteins makes the system more stable while stored and lowers its price. Among its other advantages are high selectivity (even one nucleic replacement noticeably inhibits the assembly, which eliminates the possibility of false-positive results) and flexibility (it’s possible to develop sets for any spectrum of infections). Detection of 16 pathogens at once is also what makes our system unique – there are no similar systems in Russia or the West.
How it works
Our device covers all stages of working with biological material. At the first stage, we disintegrate pathogens’ membranes (this process is called lysis) – that’s how we extract DNA or RNA. Then, the system amplifies the fragment via a chemical reaction involving salts, ferments, and other chemical substances.
Conventional methods usually involve the use of a thermal cycler to perform cycling with a constant change of temperature. We use the NASBA (nucleic acid sequence-based amplification) isothermal amplification method in a test tube heated up to a mere 41 degrees Celsius. Moreover, we have developed our own enzymes so that we wouldn’t depend on suppliers.
Then, oligonucleotides join the RNA, which has accumulated during the amplification process, form a DNA sensor, and produce components required for the visual reaction (hemin, hydrogen peroxide, and diaminobenzidine). As a result, if there was a pathogen in the sample, a reaction occurs and the test tube turns brown. If not – it remains transparent.
It’s worth mentioning how the sensors are designed. We take a pathogen – for example, E. coli – and through bioinformatics methods find the most conservative and stable site, which is the same for all E. coli. Then, in this site, we pick the optimal target for our sensors. The target will be accumulated during the amplification process. Its length is 80-160 pairs of nucleotides.
Single-stranded RNA, which machines find, is usually not linear, but folded when within the solution. This presents a problem for conventional diagnostics methods because such folded structures are very stable and self-sufficient – it’s difficult to force something external to “land” on them. However, we use this feature to our benefit, as our sensor is able to “straighten” this folded structure and identify target sites within it.
Robo-assistant

We needed to somehow implement the chemical aspect of our concept in a device and SCAMT’s Industrial Lab, headed by Alexander Vinogradov, helped us out. We were able to create a prototype of an automated micro-laboratory that consists of a robotic arm and a platform with several modules and a replaceable cartridge with reagents. For each infection, the cartridge has its own cell with sensors.
It’s important that we don’t use separate fluorophores for each pathogen, as it happens, for example, during real-time PCRs (Polymerase Chain Reactions)– otherwise, we wouldn’t have enough channels for simultaneous analysis. If the sample becomes brown as a result of the reaction, the photosensor reacts to this.
The system basically replaces a lab assistant: it moves the samples, mixes them, adds the necessary reagents, and so on. Each task has its own module: the heating one for amplification; the magnetic one for isolation and purification of DNA using magnetic particles developed at the SCAMT nanopharmaceutical laboratory; and the photosensor module for precise detection of color change.
The entire unit is placed in a container – a sealed system with integrated cleaning and disinfection functions, completely isolated from the operator. This eliminates the need for special equipment and a separate room for working with samples.
In the future, we plan to miniaturize the device and make it portable and self-powered. This project will continue for another year, so in 2021 we plan to finalize the prototype and conduct additional tests at medical institutions. Preliminary clinical studies with the participation of the Research Institute of Infections in Children have demonstrated the system’s high precision and delivered promising results.