Smart DNA strands
Although scientists are currently working on multiple gene therapy methods, they cannot offer a truly efficient cancer treatment until they figure out how to deliver a drug into a cell and enhance its efficacy and selectivity. Regular drugs do not ensure a recovery because they attack genes responsible primarily for cancer development and chemotherapy resistance. The situation may change if scientists develop treatments that will target common essential housekeeping genes that lead to apoptosis, i.e. the process of programmed cell death.
Traditional antisense oligonucleotide techniques are not good enough to suppress these types of genes as they can induce apoptosis of all body cells, thus posing a risk to human health. Nevertheless, scientists frequently refer to this long-known and simple technology in their research. It works in the following way: first, single-stranded DNA fragments find their target molecule in cells and complement it so that later the enzyme system could recognize the pair and destroy the target molecule bound with the antisense oligonucleotide.
With this method, scientists can regulate activities of various genes but not activate and deactivate the antisense agent by creating specific conditions when it is needed. Researchers from SCAMT’s Gene-Therapeutic Agents Based on Nucleic Acids Lab have further advanced the technology, resulting in the creation of a binary antisense agent. The developed agent is made up of two oligonucleotides: the first one is a therapeutic module, which tracks the target molecule for complementarity and the second one targets a tumor marker. Both parts of the chain are connected with a chemical linker, namely, a flexible chemical thread. Just like in the traditional approach, the target molecules are divided using enzymes.
“We decided to enrich the existing technique by adding two strands to track not one but two target molecules. Our binary antisense agent is activated by the marker and locates the to-be-destroyed RNA molecule in cancer cells. Thus, the technology can kill cancer cells more efficiently and not affect healthy tissues,” explains Valeriia Drozd, the first author of the paper and a second-year Master’s student at ITMO’s ChemBio Cluster.
Valeria Drozd. Photo by Dmitry Grigoryev / ITMO.NEWS
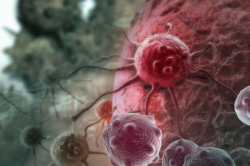
Targeting cancer cells
The researchers tested their agent in a solution with a synthetic RNase H enzyme on a fragment of green fluorescent protein (GFP) RNA and used the RNA of KRAS oncogene fragment involved in human cancer development as a tumor marker. They tried to degrade the GFP gene RNA to test the deactivation process of fluorescence. Further, the chosen RNA target will help the researchers to understand how the treatment affects cells, namely, whether the developed agent suppresses the gene in question or kills cancer cells on its own due to its toxicity.
While measuring the approach’s efficiency, the researchers took notes of cleavage percentages with and without the marker. As a result, the agent managed to destroy 9% and 53% of the molecule, respectively.
“We also used our system to identify mutations in markers. RNAs of genes that we used as markers in our study appear in both healthy and cancer cells. RNAs with mutations possess a different set of nucleotides that we can determine if we program the system to recognize a specific sequence in cells,” adds Valeriia Drozd.
The technology’s performance depends on many factors. For instance, geneticists have to successfully identify a vulnerable gene, which can trigger cell apoptosis, and find the felicitous tumor marker if they want to put this method into practice. Since this is not an easy task, the scientists are already working on their method and looking for unique genes, which can destroy all or most cancer cells when suppressed.
On top of that, the researchers still have to tackle several more challenges before they can test their method on living cells. They have to figure out how to modify the system so that it is not destroyed inside a cell following the treatment. Since any chemical modifications affect the properties of oligonucleotides, a balance needs to be maintained between the agent’s efficacy and its resistance to the human body’s protection systems. In the long-term, the team plans to develop structures based on small interfering RNAs that show greater efficiency in gene therapy.
The project is supported by the Umnik project (No. 17789ГУ/2022) and a Russian Foundation for Basic Research grant (No. 20-34-90071) within the Priority 2030 program.
Reference: Valeriia Drozd, Ahmed Eldeeb, Dmitry Kolpashchikov, Daria D. Nedorezova Binary antisense oligonucleotide (biASO) agent for cancer marker-dependent degradation of targeted RNA (Nucleic Acid Therapeutics, 2022).