What is your field of research?
I work in computational biology and mathematics in biology. A very important area of research is bioinformatics, which has existed for about 40 years but has been especially prominent the last couple of decades. The Human Genome Project, which has recently been concluded, was a major initiative in this field and prompted its rapid development. Genome assembly is, in general, a common task in bioinformatics, and by looking at the existing assembled genomes you may see what the most “popular” species are: humans, obviously; mice, which are often used for testing; and horses, because gene engineering is very important to people who work with horses and there is a lot of money in the field.
Take, for instance, one of the tasks I’m currently working on. It is well known that the influenza virus mutates from season to season, which is why it is so successful in contaminating people. If we take its genome, i.e. the sequence of the nucleotides in the DNA, we will find that in every person that we test there is a combination of various virus strains, and if we have a large enough sample of people, we can “map” the contamination and determine who contaminated who by creating a tree of the genomes and their changes. However, that requires an in-depth study of the viruses at hand.
With your education, you are more of a mathematician than a biologist. How do the two sciences coexist?
That’s right, I am more of a mathematician; I have only visited biological laboratories on tours, but I work closely with biologists. The main difficulty here is that the researchers talk in different languages, scientifically speaking. It is often said that the future is interdisciplinary, and it is true, but when it came to studying biology, it was very hard for me to find courses that I would understand and that would be interesting to me. At some point I found the course that is considered to be one of the best in the field, a course by Prof. Eric Lander from MIT, and found it much easier to understand. Midway through the course I found out that he had a PhD in cryptography and algebra, and this must have mattered a lot.

But in reality, people come to bioinformatics from both sides. A mathematician can pick up biology and a biologist can pick up mathematics. Of course, to be a good specialist you need to have a certain level of knowledge of both. A true biologist would easily notice that I come from mathematics and don’t know biology as well as they do, but that is why we collaborate and that is why it is called an interdisciplinary field. Nowadays, there are students who are already being trained to become specialists in bioinformatics; for one, ITMO has a Master’s program in Bioinformatics.
What practical tasks does bioinformatics solve? Where do the specialists work?
There is a lot of bioinformatics in pharmaceutics, for example. Sometimes it is even applied in perfumery. It is also used to develop certain diagnostic techniques. There is a startup in Moscow that aims to detect breast cancer using bioinformatics methods. In general, you can have your DNA studied and sequenced to have specialists tell you the probability of you developing certain diseases. For some diseases, this process is already well established and even trivial in some cases.
The abovementioned building of the virus trees can sometimes be used to solve legal cases (who contaminated who) and there have been precedents for this in Europe and the US. Also, this can help us find the point in the mutation of the virus at which it becomes more virulent, as in better at infecting and damaging its host.

Another application for bioinformatics is agriculture, which is an important sector of the economy. Why do shrimps die? How do we edit the genes of produce to make it more resilient to transportation? Bioinformatics might answer some of these questions.
Forensic science is very well known to people from various movies and TV series. Specialists can find the culprit using the DNA fingerprint from any cell left on the crime scene. There is even a project in the USA that helps acquit people wrongly sentenced to death or life imprisonment using this technology.
Another example of the implementation of bioinformatics is the company 23andMe that allows you to study your DNA to find your ancestry all around the world.
What fields of mathematics are important for bioinformatics?
In bioinformatics, graph theory plays a key role. Building evolutionary trees falls under graph theory, collecting genomes uses graph theory, and the folding of proteins also implements graph theory to a certain extent. Since folding protein is all about their 3D shape, it also involves some geometry. In general, continuous mathematics (what some people with no background in math would call simply mathematics) is taught more in universities, but biology requires a lot of what’s called discrete mathematics, which is mathematics of discrete values, and graph theory is a part of it.
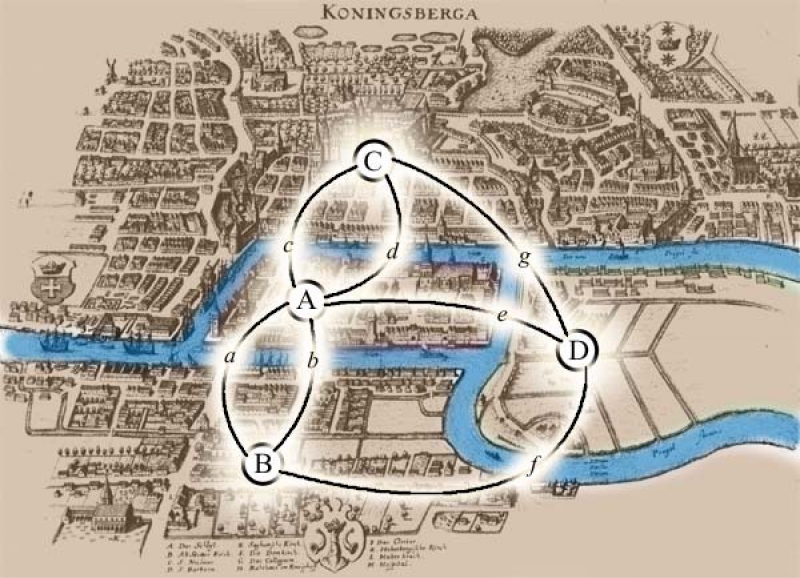
Graph theory is actually very old: there is the famous Seven Bridges of Königsberg problem; Euler trying to walk all the bridges without stepping on the same island twice is actually relevant to the study of genomes. In a way, you could say that mathematics had been preparing for bioinformatics all along.
And of course there is a lot of statistics, probability theory and combinatorics.
What are some current issues that bioinformatics tackles?
Reading a genome is still a challenging task sometimes. Humans are one thing, as we have spent 10 years collecting our genome, but corn (the genome of which is larger than that of humans) is another thing and soil is a different thing altogether, because of the amount of living organisms it has.
A very important general task is determining the phenotype using the genotype. It has been done for some specific cases: we know what some genes stand for and what they change in the appearance of the organisms, such as their sex, eye color, hair color, and the risk of certain diseases, but there is still a lot to learn.
There are many tasks in immunology that revolve around understanding how the immune system works and how cells learn to fight foreign organisms. Of course, a huge amount of resources goes into finding a cure for cancer, and bioinformatics plays an important role here, too, since cancer is a genetic disease. There are many challenges ahead of us and biology in general has changed completely over the last 50 years, which is why this science is so interesting.

You have studied and worked in the USA. What drew you there?
During my Master’s studies at St. Petersburg State University, as well as when I worked on my PhD thesis, I traveled a lot to Germany and worked there as a mathematician. In 2010, I returned to Russia. Back then I wasn’t studying bioinformatics; instead, I was working with random matrices and only at the end of my PhD studies I started getting interested in bioinformatics.
At that point I was looking for places to work and the US seemed like a reasonable choice to me. I think that everyone should visit the US at some point in their life. There was a person there who worked in the same field as I did; when I contacted him, I simply wanted to collaborate on something, but it turned out that he had an open postdoc position.
Washington, DC is a very unusual city that feels artificial in certain ways. There are only government officials there and everything is suited to their needs. In general, American cities are very different from the ones in Europe and Russia, as well, and you need to have a car to get around. For example, you can’t really walk around Los Angeles; you have to travel by car between the sights, shops, bars, etc. The same can be said about Atlanta. Americans live in low-rise buildings and therefore a large mall that provides services to the whole neighborhood will be located far away from a lot of the houses. But Washington is much more European in that sense, it’s more compact. A very political city.
Obviously, it didn’t really concern me; I was working at a university. And there are a lot of museums around, which I found great.
Why did you choose ITMO University?

At some point I got weary of the US and decided that I want to return to Europe. And Europe turned out to be ITMO University. In a lot of ways ITMO’s offer was unique, as it felt good to return to speaking my mother tongue again, and the students here are well-educated. I think the students were the main reason that I stayed here. Currently, I work with a lot of students and coordinate a number of projects. I would say that the students here are very good at math and I find working with them very comfortable. I continue to collaborate with researchers in the US, which gives me access to data from a number of hospitals, and here I have talented students who can work with that data.
I like it here at ITMO. My current plans are to continue working on projects and to continue working with students.




